SMRU QC config file structure
All ArgosQC workflows are structured using a JSON config file. The
config file is hierarchical in structure with as many as 4 blocks:
setup, harvest, model, and
meta. The meta block is only required if no
metadata file is provided in the setup block. These files
can be constructed manually or programmatically and provide the ability
to define certain aspects of the QC workflow. Below is the config file
for an IMOS QC workflow on SMRU GPS SRDL-CTD tags deployed on olive
ridley turtles on the Tiwi Islands, Australia. All SMRU tag data are
organised by deployment campaign ids, e.g., ct188. A
separate QC workflow (config file) must be generated for each deployment
campaign.
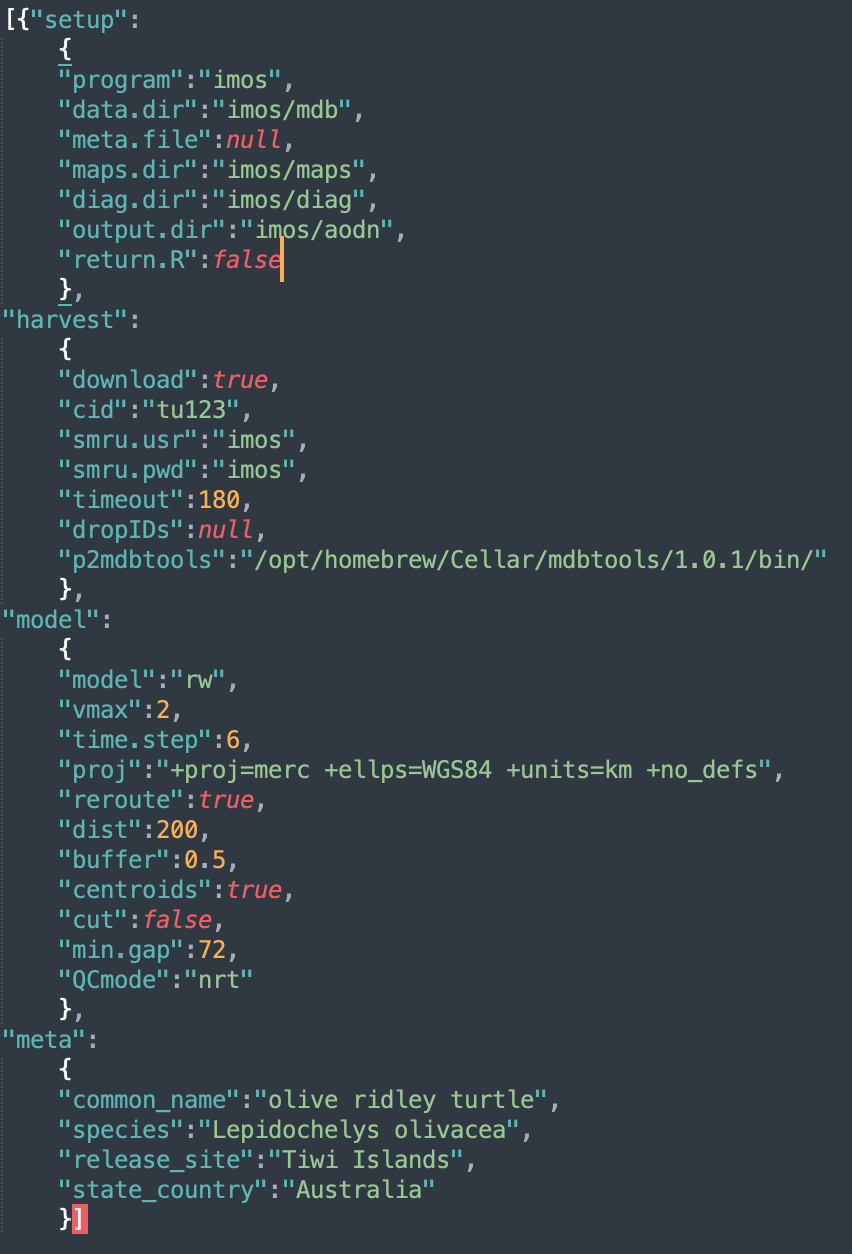
The block-specific parameters are as follows:
-
setupconfig block specifies the program overseeing data assembly & paths to required data, metadata & output directories:-
programthe national (or other) program of which the data is a part. Current options are:imosoratn. -
data.dirthe name of the data directory. Must reside within thewd. The directory will be created if it doesn’t exist. -
meta.filethe metadata filename. Must reside within thewd. Can be NULL, in which case, themetaconfig block (see below) must be present & tag-specific metadata are scraped from the SMRU data server (provided SMRU server access credentials are supplied - seeharvestblock, below). -
maps.dirthe directory path to write diagnostic maps of QC’d tracks. -
diag.dirthe directory path to write diagnostic time-series plots of QC’d lon & lat variables. -
output.dirthe directory path to write QC output CSV files. Must reside within thewd. -
return.Ra logical indicating whether the function should return a list of QC-generated, internal R objects. This results in a single large object returned to the R work space containing the following elements:-
cidthe SMRU campaign ID -
dropIDsthe SMRU Reference ID’s dropped from the QC process -
smruthe SMRU tag data tables extracted from the downloaded .mdb file -
metathe working metadata -
locs_sfthe projected location data to be passed as input to the SSM -
fit1the initial SSM output fit object -
fit2the final SSM output fit object including re-routed locations if specified. -
smru_ssmthe SSM-annotated SMRU tag data tables. This output object can be useful for troubleshooting undesirable results during delayed-mode (supervised) QC workflows.
-
-
-
harvestconfig block specifies data harvesting parameters:-
downloada logical indicating whether tag data are to be downloaded from the SMRU data server or read from the localdata.dir. -
cidSMRU campaign ID. -
smru.usrSMRU data server username as a string. -
smru.pwdSMRU data server password as a string. -
timeoutextends the download timeout period a specified number of seconds for slower internet connections. -
dropIDsthe SMRU ref ID’s that are to be ignored during the QC process. Can be NULL. -
p2mdbtools(optional) provides the path to the mdbtools library (required extract data from.mdb-Microsoft Access Database files) if it is installed in a non-standard location (e.g., on Macs when installed via Homebrew). Can be set to NULL otherwise.
-
-
modelconfig block specifies model- and data-specific parameters:-
modelthe aniMotum SSM model to be used for the location QC - typically eitherrworcrw. -
vmaxfor SSM fitting; max travel rate (m/s) to identify implausible locations -
time.stepthe prediction interval (in decimal hours) to be used by the SSM -
projthe proj4string to be used for the location data & for the SSM-estimated locations. Can be NULL, which will result in one of 5 projections being used, depending on whether the centroid of the observed latitudes lies in N or S polar regions, temperate or equatorial regions, or if tracks straddle (or lie close to) -180,180 longitude. -
reroutea logical; whether QC’d tracks should be re-routed off of land (default is FALSE). Note, in some circumstances this can substantially increase processing time. Default land polygon data are sourced from theropensci/rnaturalearthhiresR package. -
distthe distance in km from outside the convex hull of observed locations from which to select land polygon data for re-routing. Ignored ifreroute = FALSE. -
barrieran optional filepath to an alternate polygon data file (shapefile) for the land (or other) barrier. For example, higher resolution local coastline data can be supplied, provided the data extend at leastdistkm beyond the extent of the track data. -
bufferthe distance in km to buffer rerouted locations from the coastline. Ignored ifreroute = FALSE. -
centroidswhether centroids are to be included in the visibility graph mesh used by the rerouting algorithm. See?pathroutr::prt_visgraphfor details. Ignored ifreroute = FALSE. -
cutlogical; should predicted locations be dropped if they lie within in a large data gap (default is FALSE). -
min.gapthe minimum data gap duration (h) to be used for cutting predicted locations (default is 72 h) -
QCmodeone of eithernrtfor Near Real-Time QC ordmfor Delayed Mode QC.
-
-
metaconfig block specifies species and deployment location information. This config block is only necessary when no metadata file is provided in thesetupblock.-
common_namethe species common name (e.g., “southern elephant seal”) -
speciesthe species scientific name (e.g., “Mirounga leonina”) -
release_sitethe location where tags were deployed (e.g., “Iles Kerguelen”) -
state_countrythe country/territory name (e.g., “French Overseas Territory”)
-
With a completed config file, the standard call to initiate the QC workflow within R is:
smru_qc(wd = "test", config = "imos_config.json")where wd is the file path for the working directory
within which all QC data/metadata inputs are downloaded (or read) and
outputs are written.
Additional details on config parameters
The proj argument specifies the projection (as a
proj4string) to which the tag-measured locations are
converted as input to the QC state-space model (SSM), ie. the working
projection in km for the SSM. Any valid
proj4string may be used, provided the units are in
km. If proj is left as NULL then
the QC algorithm will project the data differently depending on the
centroid latitude of the tracks. The default projections are:
| Central Latitude or Longitude | Projection (with +units=km) |
|---|---|
| -55 to -25 or 25 to 55 Lat | Equidistant Conic with standard parallels at the tracks’ 25th & 75 percentile Latitudes |
| < -55 or > 55 Lat | Stereographic with origin at the tracks’ centroid |
| -25 to 25 Lat | Mercator with origin at the tracks’ centroid |
| -25 to 25 Lat & Long straddles -180,180 | Longitudes are shifted to 0, 360 and a Mercator with origin at tracks’ centroid |
The model argument specifies the aniMotum
SSM to be used; typically either rw or crw.
The latter is usually less biased when data gaps are absent, the former
is best when data gaps are present. A general recommendation is to use
model:rw as the SSM for unsupervised (e.g.,
NRT) QC workflows. The SSM fitting algorithm has a few fundamental
parameters that need to be specified; vmax is the animals’
maximum plausible travel rate in
ms.
For example, vmax:3 is usually appropriate for
seals and vmax:2 for turtles. The SSM
prediction interval in hours is specified with time.step.
Decimal hours can be used for time.steps shorter than 1 hr.
This time interval determines the temporal resolution of the predicted
track. The predicted track locations provide the basis for interpolation
to the time of each tag-measured ocean observation or behavioural event.
Typically, 6 hours is appropriate for most Argos data collected from
seals and turtles but a finer time interval may be required for faster
moving species and/or more frequently measured ocean observations, and a
coarser interval for more sporadically observed locations. Further
details on SSM fitting to Argos and GPS data are provided in the
associated R package aniMotum vignettes and
in Jonsen
et al. 2023.
When animals pass close to land some SSM-predicted locations may
implausibly lie on land. Often, this is due to the spatial and temporal
resolution of the Argos tracking data. In these cases, SSM-predicted
locations can be adjusted minimally off of land by setting
reroute:true. The pathroutr R
package is used for efficient rerouting. In this case, additional
arguments should be specified:
dist - the distance in km beyond track locations from
which coastline polygon data should be sampled (smaller provides less
information for path re-routing, greater increase computation time)
barrier - an optional parameter that can provide an
alternate spatial polygon dataset , as a shapefile, for the land (or
other barriers to movement). Typically, this alternate dataset would be
a localised, high-resolution coastline dataset.
buffer - the distance in km to buffer rerouted locations
from the coastline
centroid- whether to include the visibility graph
centroids for greater resolution
SSM-predicted tracks can be cut
(cut:true) in regions where large location
data gaps exist. These location data gaps can occur when the tags are
unable to transmit for extended periods or when animal surfacing occurs
during periods of Argos satellite unavailability (more common closer to
the equator than at higher latitudes). In this case,
min.gap is used to specify the minimum data gap duration
(h) from which to cut SSM-predicted locations. This will limit
interpolation artefacts due to implausible SSM-predicted locations in
excessively long data gap periods.
The QCmode sets whether the QC is being conducted in
delayed-mode dm or near real-time nrt.
Delayed-mode is reserved for when tag deployments have ended and usually
involve greater user intervention; such as making decisions on removing
aberrant portions of a deployment (e.g., as tag batteries begin
failing). The nrt mode is mean to be fully automated and
only used while a deployment is active. In both cases, the output .CSV
and plot file names will include the QCmode as a
suffix.