SMRU SRDL tag QC workflow
The first step to initiate any ArgosQC workflow is to
construct a JSON config file (see SMRU_config_file).
The SMRU SRDL tag QC workflow has a number of data/metadata processing, model fitting, data file annotation and output steps. Each of these steps is encapsulated in an ArgosQC function:
Table 1. SMRU SRDL tag QC workflow functions listed in order of
operation. For standard near real-time (NRT) QC workflows, these
functions are implemented via the wrapper function,
smru_qc(). For examples of how to implement the individual
functions refer to the R help pages, e.g., type
?download_data in the R console.
| SMRU QC function | Description |
|---|---|
download_data |
Downloads tag data .mdb file from SMRU server &
writes to the specified data.dir. |
smru_pull_tables |
Extracts tag data files from the downloaded .mdb file
to a named list object in the R workspace, See
?smru_pull_tables for an example. |
get_metadata |
Either loads deployment metadata from a CSV file
specified in the config file setup block, or
builds the metadata from the tag manufacturer’s data portal in
combination with species and deployment site attributes provided in the
config file meta block. |
smru_prep_loc |
Prepares location data for SSM fitting by
restructuring the SMRU diag & gps (if present) tag data files,
truncating start and end dates (for NRT QC) based on dates of first
& last CTD profile, projecting locations from lon-lat to the
config file specified proj4string. |
multi_filter |
Applies a 1st pass of the SSM model using parameters
specified in the config file model block.
SSM’s are fit in parallel across n available processors. |
redo_multi_filter |
Applies a 2nd pass to refit the SSM to any tag
location datasets that failed to converge on the first pass. Uses
automatically revised parameters to help ensure convergence. Reroutes
any locations off of land, if model:reroute:true in
config file. |
ssm_mark_gaps |
Identifies & marks SSM predicted & rerouted locations in track segments with data gaps of a specified minimum duration. Typically used in DM QC’s only. |
smru_append_ssm |
Appends SMRU tag data files with SSM-derived coordinates & uncertainty (lon,lat,x,y, x.se, y.se) for each record. |
smru_clean_diag |
Restructures the diag file for diagnostic plots |
diagnostics |
Generates a map of all QC’d tracks & time-series plots of longitude & latitude for quick assessment of SSM fit to the tag location datasets. |
smru_write_csv |
Combines QC-annotated tag data files across individual tags, applies tests for expected variables types and ranges (IMOS only for now), writes data files to CSV as final QC outputs |
If a more complicated NRT workflow is required, e.g., with custom processing in between the standard workflow steps, the functions can be called separately from an R script so that intermediate results can be checked. This is also the recommended approach for all delayed-mode (supervised) QC workflows.
In this example, the main QC outputs were written to .CSV files in
the specified output directory, output.dir. Each .CSV file
includes the name of the SMRU data table, when present
(ctd, diag, dive,
haulout, summary) or the QC file
(metadata, ssmoutputs). For QC workflows with
ATN data, each of these file names is appended with the species’
AnimalAphiaID and the ADRProjectID. For IMOS
and other programs, the file names are appended with the SMRU campaign
ID (e.g., ct182).
The diag files show the SSM fit (red) overlaid on the tag-measured
Argos &/or GPS locations (blue). The dark grey vertical bars denote
the time period tags were actively recording locations but the seal(s)
either had not yet gone to sea (no recorded diving activity - left
side), or the CTD sensor had failed (e.g., grey bar on right side of
tu123-Catherine-25). By default, the QC model does not fit
to data in these time periods. These plots help judge whether the SSM
fits have artefacts that need addressing - typically only addressed
during a delayed-mode QC workflow. 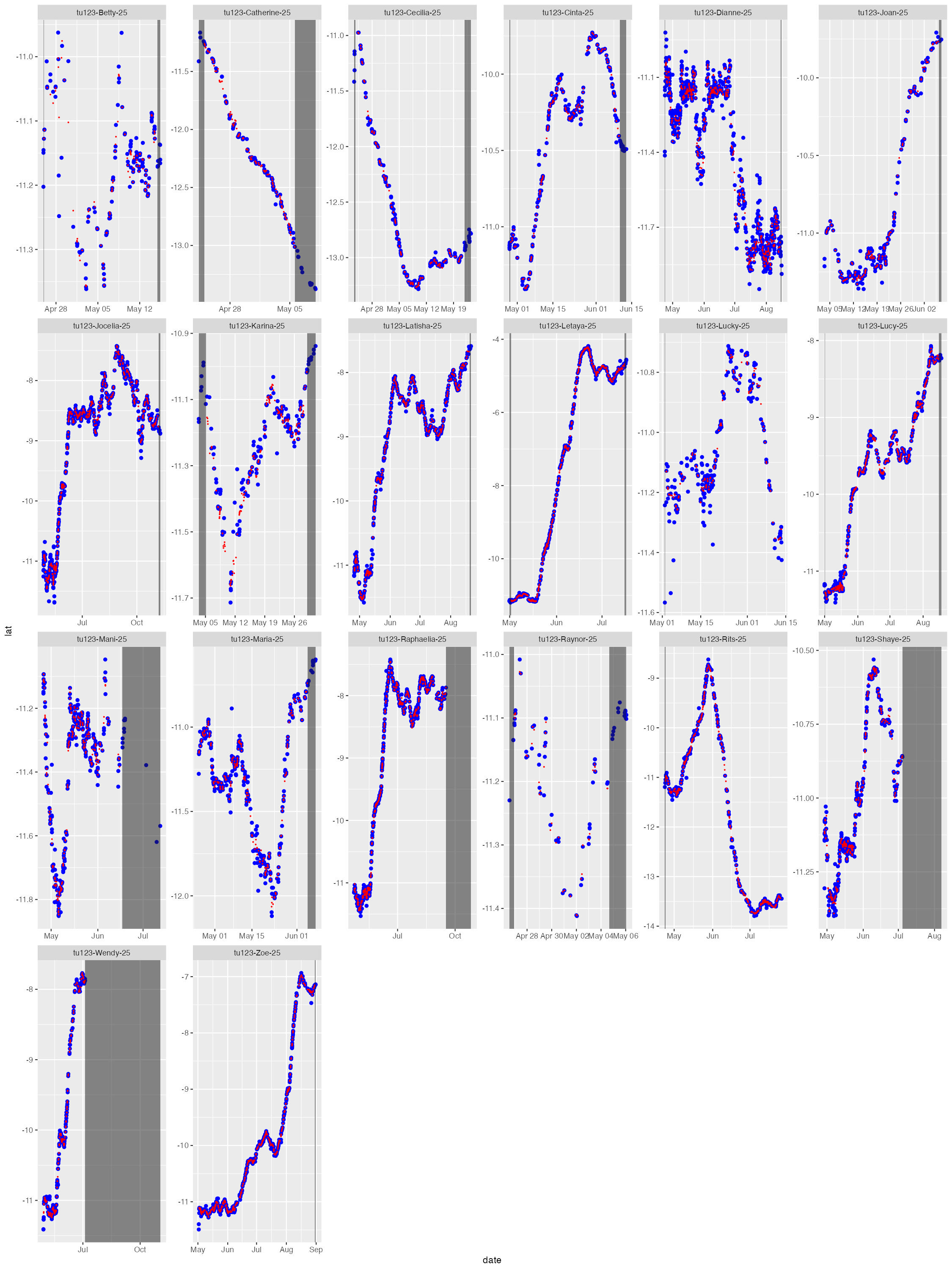
The map file shows the SSM-predicted tracks (blue) and current last
estimated location (red) for each deployed tag. The map files are
annotated by the QC date so they are not overwritten by successive QC
runs. 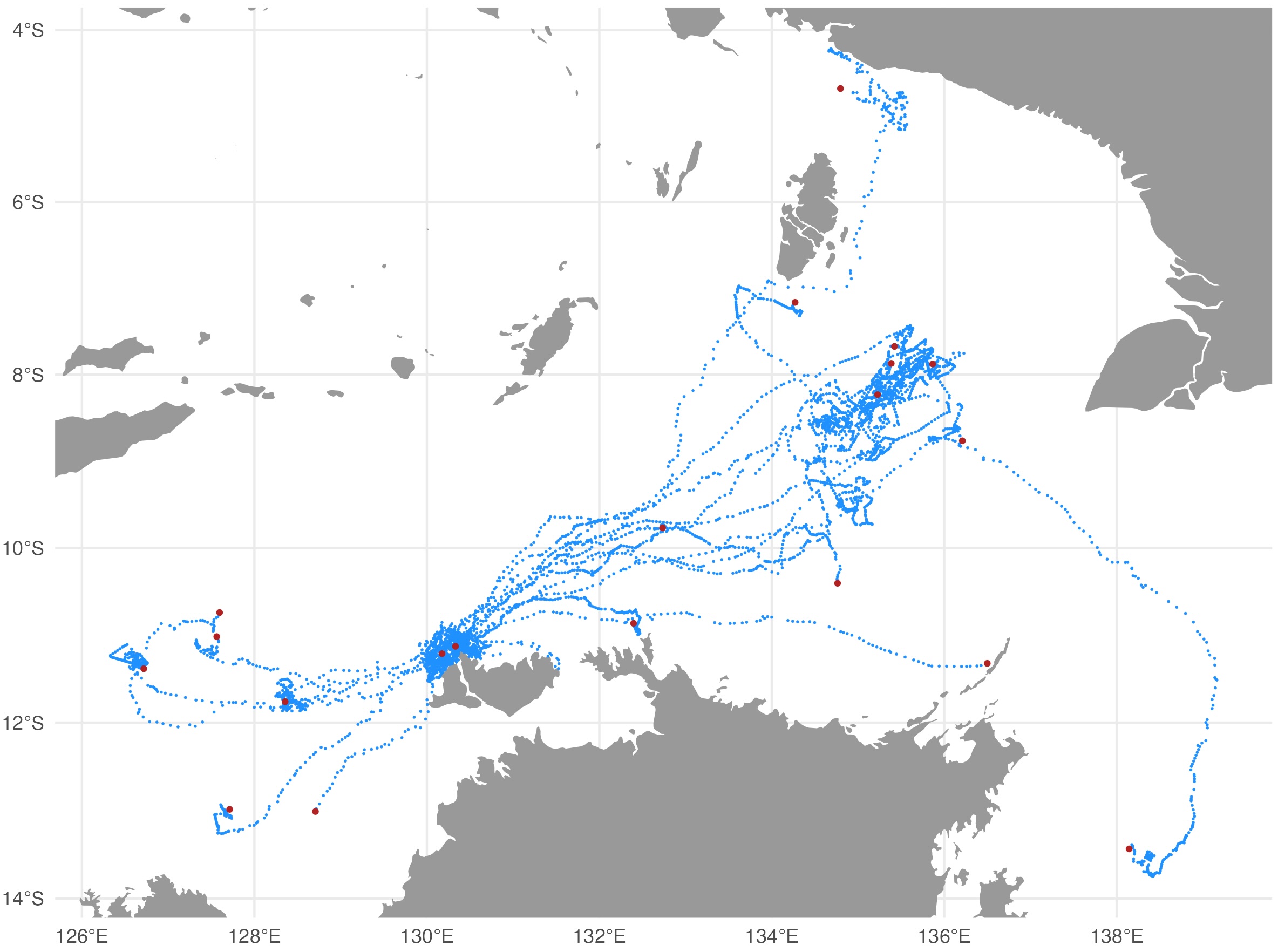
Output .CSV files
The QC’s main outputs, the .CSV files contain all records from the
original SMRU data tables and are appended with the following additional
columns: ssm_lat, ssm_lon, ssm_x,
ssm_y, ssm_x_se, ssm_y_se. These
are the QC’d locations and their uncertainty estimates interpolated to
the time of each record. The ssm_x, ssm_y
variables are the coordinates from the QC workflow projection (in km)
and ssm_x_se, ssm_y_se are the associated
standard errors (in km). Note that NA’s may be present in the
QC-appended location variables, particularly at the start and/or end of
individual tracks. This is typically indicative of track portions prior
to animals going to sea (at deployment start) and portions when either
the CTD or pressure sensor failed, eg. due to biofouling or seawater
ingress, but tag still transmitted locations (near deployment end).
Metadata .CSV file
If an input deployment/tag metadata file is provided then the output metadata file contains all the original metadata records plus the following variables describing the QC workflow applied to the data:
-
qc_start_date- the track datetime (UTC) at which the QC workflow was started. -
qc_end_date- the track datetime (UTC) at which the QC workflow was ended. -
qc_proj4string- the projection used for QC’ing the locations, as a proj4string. -
qc_method- denotes theArgosQCR package was used. -
qc_version- denotes the version number of theArgosQCR package used. -
qc_run_date- the datetime (UTC) when the QC was applied to the data.
Note, these variables are not appended to the metadata for IMOS QC workflows due to IMOS - AODN metadata specifications.
SSMOutputs .CSV file
The SSMOutputs file contains the SSM-predicted locations at the
time.step specified prediction interval. The time of the
first location is set to the time of the first tag-measured location
passed to the model. This may or may not be the first tag-measured
location in the tag datafile, depending on whether the animal-borne tag
was immediately at sea. The location coordinates are provided as:
lon, lat, x, y, and
location uncertainty as x_se, y_se. The planar
coordinates and uncertainty estimates always have units in km. Their
coordinate projection is provided in the metadata .CSV file
(qc_proj4string).