Wildlife Computers (WC) tag QC workflow
The first step to initiate any ArgosQC workflow is to
construct a JSON config file (see WC_config_file).
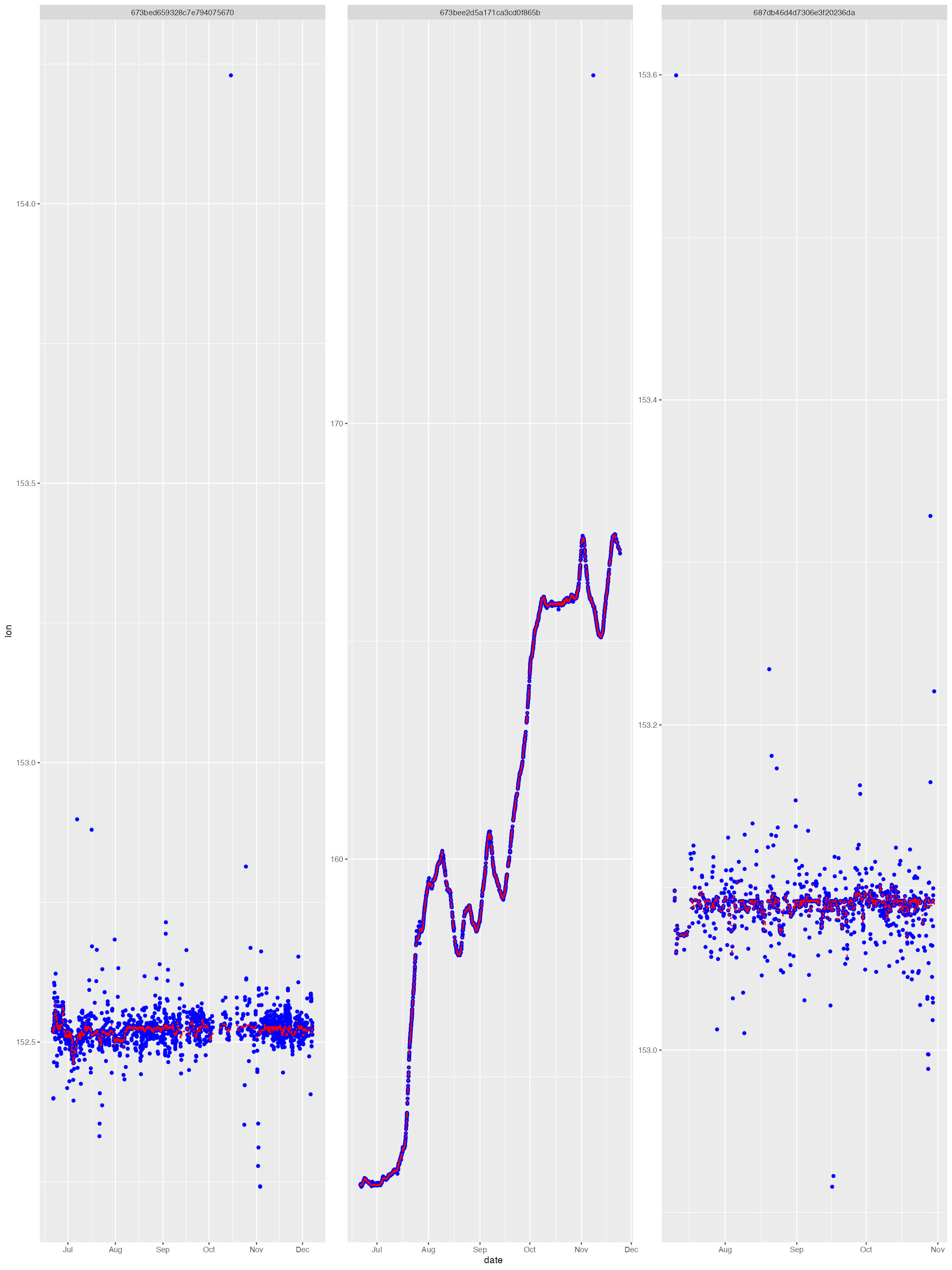
The diag files show the SSM fit (red) overlaid on the tag-measured Argos &/or GPS locations (blue). The dark grey vertical bars denote the time period tags were actively recording locations but the seal(s) had not yet gone to sea (no recorded diving activity). By default, the QC model does not fit to data in these time periods. These plots help judge whether the SSM fits have artefacts that need addressing - typically only addressed during a delayed-mode QC workflow.
The map file shows the SSM-predicted tracks (blue) and current last
estimated location (red) for each deployed tag. The map files are
annotated by the QC date so they are not overwritten by successive QC
runs. 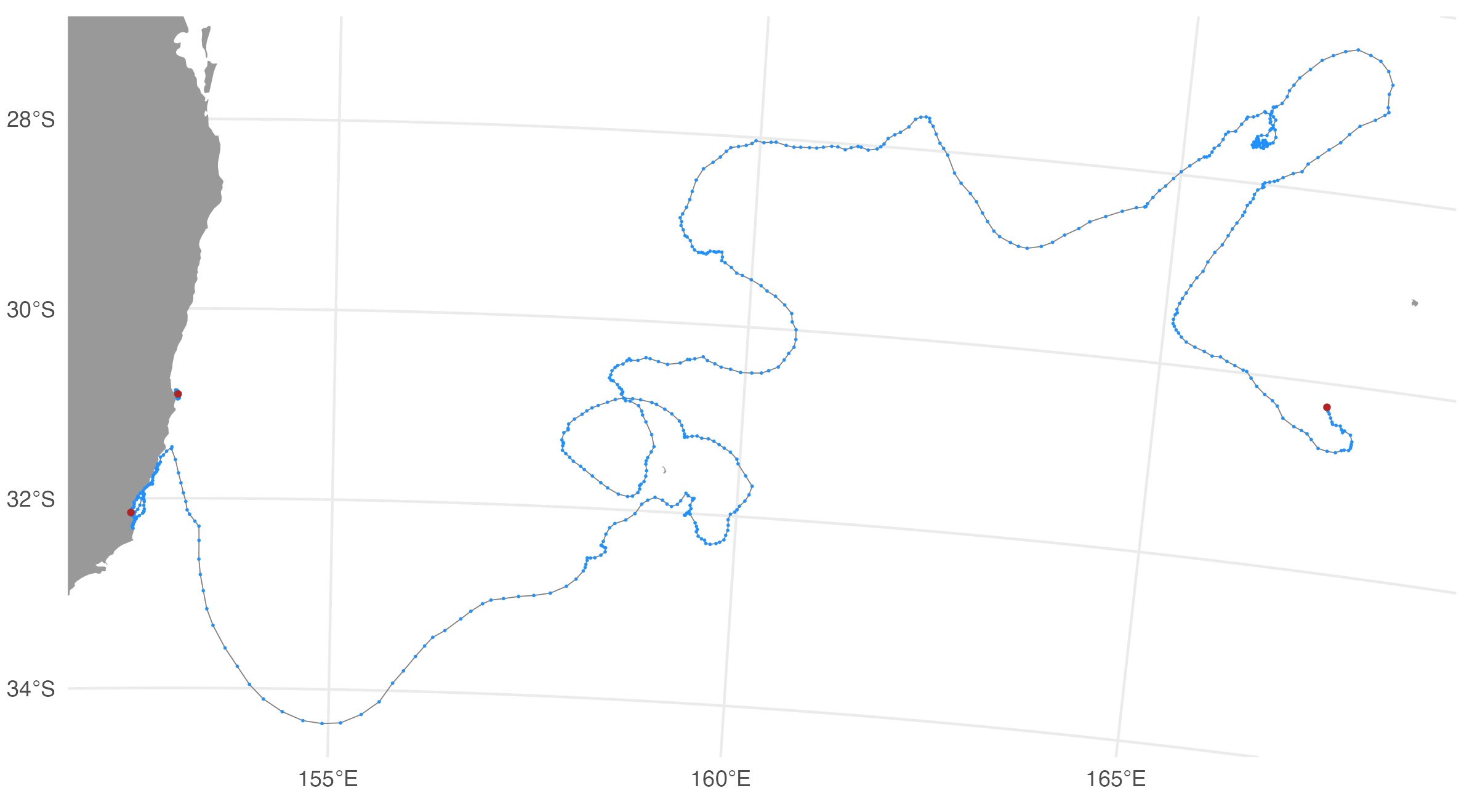
Output .CSV files
The QC’s main outputs are the Wildlife Computers tag data files
appended with the same QC’d location variables as the SMRU tag QC output
files: ssm_lat, ssm_lon, ssm_x,
ssm_y, ssm_x_se, ssm_y_se. The
output .CSV files necessarily depend on the specific type of Wildlife
Computers tag(s) that is/are being QC’d. Currently, the
ArgosQC workflow accommodates SPOT, SPLASH and SCOUT tag
data. Combined across these tag types, the following output .CSV files
(per Wildlife Computers) are provided:
DSAECDHistosFastGPSHistosLocationsMinMaxDepthMixLayerPDTsSST
Where one of these files can have a data structure that differs
between specific tag types, the file names are appended with the tag
type. For example, the ECDHistos file structure differs
between SCOUT DSA and SCOUT TEMP 361A tags, so these file names appear
as either ECDHistos_SCOUT_DSA or
ECDHistos_SCOUT_TEMP_361A. Similar scenarios for other tag
types/versions will be incorporated as required in future versions of
ArgosQC.
In the generic metadata case, ie. for all programs other than ATN,
each of the WC QC output file names are appended with the species’
common name and the QCmode suffix - _nrt or
_dm. For ATN data and metadata, each file is appended with
the AnimalAphiaID, the ATN ADRProjectID and the QCmode
suffix - _nrt or _dm.
Metadata .CSV file
If an input deployment metadata file is provided then the output metadata file contains all the original metadata records plus the following variables describing the QC workflow applied to the data:
-
qc_start_date- the track datetime (UTC) at which the QC workflow was started. -
qc_end_date- the track datetime (UTC) at which the QC workflow was ended. -
qc_proj4string- the projection used for QC’ing the locations, as a proj4string. -
qc_method- denotes theArgosQCR package was used. -
qc_version- denotes the version number of theArgosQCR package used. -
qc_run_date- the datetime (UTC) when the QC was applied to the data.
ssmoutputs .CSV file
The ssmoutputs file contains the SSM-predicted locations at the
pred.int specified prediction interval. The time of the
first location is set to the time of the first tag-measured location
passed to the model. This may or may not be the first tag-measured
location in the tag locations data file, depending on whether the
animal-borne tag was immediately at sea. The location coordinates are
provided as: lon, lat, x,
y, and location uncertainty as x_se,
y_se. The planar coordinates and uncertainty estimates
always have units in km. Their coordinate projection is provided in the
metadata .CSV file (qc_proj4string).